Note
This page is a reference documentation. It only explains the function signature, and not how to use it. Please refer to the user guide for the big picture.
nidl.metrics.alignment_score¶
- nidl.metrics.alignment_score(z1, z2, normalize=True, alpha=2, eps=1e-12)[source]¶
Compute the alignment score between two embeddings [1].
This metric measures how closely aligned two embeddings
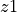 and
and
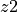 . It corresponds to the expected powered Euclidean distance
between embeddings. Lower values = better alignment.
. It corresponds to the expected powered Euclidean distance
between embeddings. Lower values = better alignment.Formally:
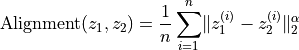
with
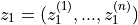 and
and
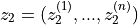
- Parameters:
- z1: torch.Tensor or np.ndarray, shape (n_samples, n_features)
Embeddings from the first view / augmentation.
- z2: torch.Tensor or np.ndarray, shape (n_samples, n_features)
Embeddings from the second view / augmentation. Must have the same shape as z1.
- normalize: bool, default=True
If True, each vector is L2-normalized before computing the alignment, as done in contrastive methods that operate on the unit hypersphere (SimCLR, MoCo, etc.).
- alpha: int or float, default=2
Exponent applied to the Euclidean distance. - alpha=2 corresponds to the original definition in the paper. - alpha=1 gives average L2 distance.
- eps: float, default=1e-12
Small value added to avoid division by zero.
- Returns:
- scoretorch.Tensor or numpy scalar
The alignment score. Lower is better.
If inputs are tensors → returns a 0-dim torch.Tensor.
If inputs are NumPy arrays → returns a numpy.float64.
References
[1]T. Wang, P. Isola, “Understanding Contrastive Representation Learning through Alignment and Uniformity on the Hypersphere”, ICML 2020.
Examples using nidl.metrics.alignment_score¶

Visualization of metrics during training of PyTorch-Lightning models